Page Content (OCR)
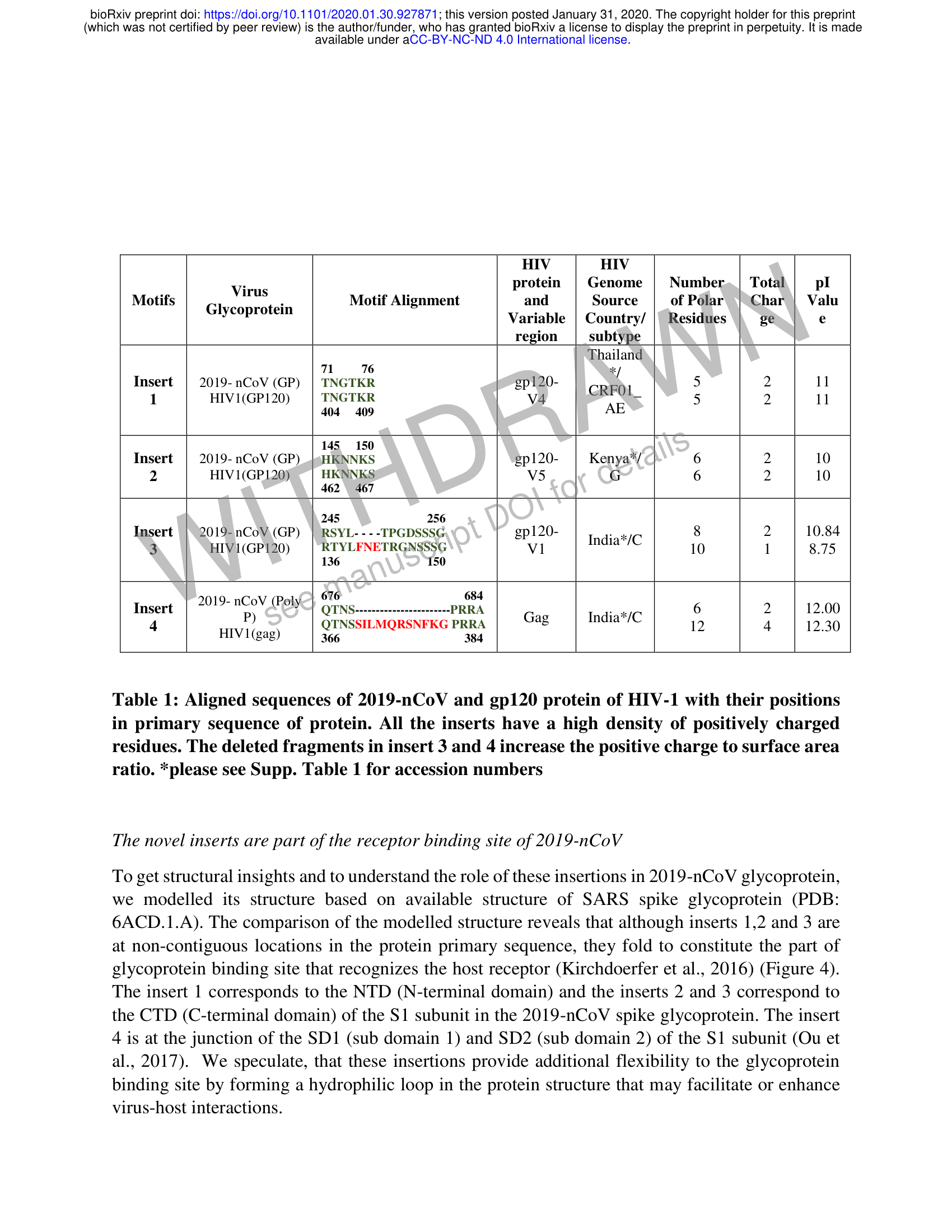
bioRxiv preprint doi: https://doi.org/10.1101/2020.01.30.927871; this version posted January 31, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. HIV HIV protein | Genome | Number,.| Total pl Motifs Gl virus on Motif Alignment and Source of Polar. | Char | Valu yeop Variable | Country/ || Residues ge e region subtype Thailand 711 (76 */ Insert 2019-nCoV (GP) | TNGTKR gp120- CRFOL 5 2 11 1 HIV1(GP120) TNGTKR v4 = 5 2 rT 404 409 AE 145 150 Insert | 2019-nCoV (GP) _.|sHKNNKS gp120- Kenya*/ 6 2 10 2 HIV1(GP120)) | HKNNKS V5 G 6 2 10 462 467 245 256 Insert | 2019-nCoV (GP) | RSYL-- --TPGDSSSG gp120- India*/C 8 2 10.84 3 HIVI(GP120) | RTYLFNETRGNSSSG V1 10 1 8.75 136 150 676 684 Insert | 2019 ae POW) Op NS aannnnnnnseeneeeneeee PRRA G India*/C 6 2 12.00 4 ) QTNSSILMQRSNFKG PRRA ag nda 12 4 12.30 HIV 1 (gag) 366 384 Virus Glycoprotein Motif Alignment 11 11 10 10 10 12 Gag Table 1: Aligned sequences of 2019-nCoV and gp120 protein of HIV-1 with their positions in primary sequence of protein. All the inserts have a high density of positively charged residues. The deleted fragments in insert 3 and 4 increase the positive charge to surface area ratio. *please see Supp. Table 1 for accession numbers To get structural insights and to understand the role of these insertions in 2019-nCoV glycoprotein, we modelled its structure based on available structure of SARS spike glycoprotein (PDB: 6ACD.1.A). The comparison of the modelled structure reveals that although inserts 1,2 and 3 are at non-contiguous locations in the protein primary sequence, they fold to constitute the part of glycoprotein binding site that recognizes the host receptor (Kirchdoerfer et al., 2016) (Figure 4). The insert 1 corresponds to the NTD (N-terminal domain) and the inserts 2 and 3 correspond to the CTD (C-terminal domain) of the S1 subunit in the 2019-nCoV spike glycoprotein. The insert 4 is at the junction of the SD1 (sub domain 1) and SD2 (sub domain 2) of the S1 subunit (Ou et al., 2017). We speculate, that these insertions provide additional flexibility to the glycoprotein binding site by forming a hydrophilic loop in the protein structure that may facilitate or enhance virus-host interactions. The novel inserts are part of the receptor binding site of 2019-nCoV